
吳志航 (Wu, Chih-Hang)
副研究員
- 英國東安格利亞大學生物學系 博士
- Plant Immunity; Molecular Plant-microbe Interactions
- wuchh@gate.sinica.edu.tw
- wuchh@as.edu.tw
- +886-2-2787-1142 (Lab: )
- +886-2-2787-1141 (Office: R420)
- Lab Website
- Academia Sinica Archive
- ORCID
- Web of Science (WOS)
- Google Scholar
植物先天免疫系統
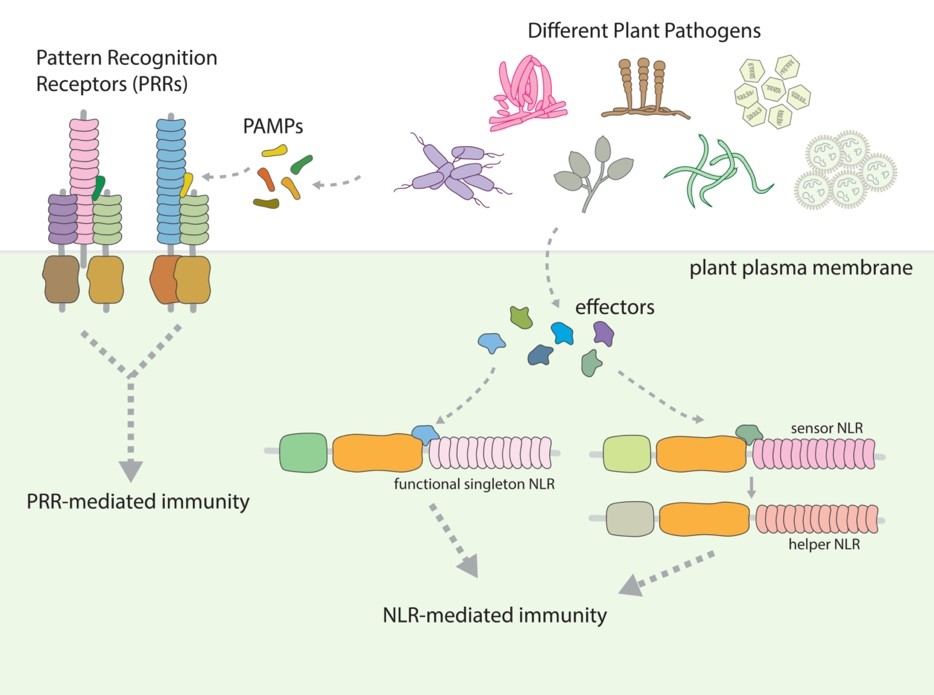
植物在生長過程中會和許多微生物有接觸,其中有許多微生物能夠感染植物並造成病害。有一些植物病原性的微生物能夠感染重要的經濟作物,造成農業上重大的損失。為了抵抗植物病原菌的侵染,植物利用其免疫系統來偵測病原菌並限制病原菌的生長。其中包含利用細胞表面的PRR免疫受體及細胞內部的NLR 免疫受體來辨識來自病原菌的分子並引發免疫訊息傳導。許多的免疫受體具有抗病蛋白的功能,能夠保護植物避免病原菌的感染,因此非常具有在農業上應用的價值。
PRR位於細胞膜上,偵測胞外來自病原菌的PAMP,而NLR位於細胞內部,偵測由病原菌分泌到細胞內部的效應蛋白。在偵測到來自病原菌的分子之後,這些免疫受體活化下游防禦訊息傳遞,達成由PRR介導的免疫反應或由NLR介導的免疫反應。有些NLR可以單獨作用,不需依賴其他NLR來辨識病原菌及引發防禦反應。有些NLR則需共同作用,這些共同作用的NLR可以進一步地被分為可以辨識病原菌的“偵測型NLR”及引發下游免疫訊號的“輔助型 NLR”。
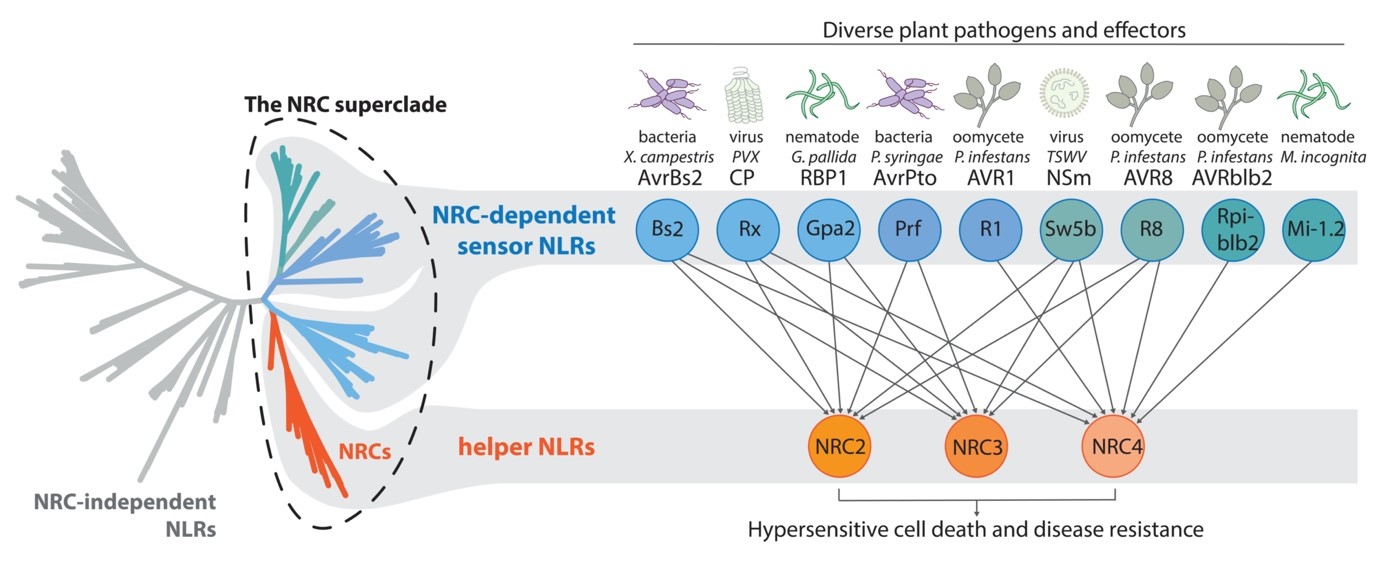
茄科植物的NRC免疫受體網絡
近期的研究發現NLR可以單獨作用、成對作用或是形成複雜的網絡。茄科植物的NRC網絡由多個能夠辨識不同病原菌的偵測型NLR及三個主要的輔助型 NLR (NRC2, NRC3, 及NRC4)所組成。這三個NRC在功能上有部分的重疊性但又對不同的偵測型NLR具有專一性。除此之外,NRC家族和依賴NRC的偵測型NLR在演化樹上屬於同一個大演化支內的不同次分支群。這些研究不僅顯示NLR免疫受體可以形成複雜的網絡來達成對於多種不同病原體的抗病性,更將NLR免疫受體的演化進程和涉及的免疫訊息傳遞進行連結。
本實驗室的研究著重於了解植物免疫系統的演化及功能,我們希望能夠回答關於NRC網絡的以下三個問題:
- 輔助型 NLR與偵測型NLR如何協同作用?
- NRC網絡如何在不同植物組織有專一化的現象?
- NRC網絡在不同的植物演化支系內有什麼不同的演化進程?
- Selvaraj M, Toghani AA, Pai H, Sugihara Y, Kourelis J, Yuen ELH, Ibrahim T, Zhao H, Xie R, Maqbool A, De la Concepcion JC, Banfield MJ, Derevnina L, Petre B, Lawson DM, Bozkurt TO, Wu CH, Kamoun S and Contreras MP. 2024. Activation of plant immunity through conversion of a helper NLR homodimer into a resistosome. PLOS Biologyhttps://doi.org/10.1371/journal.pbio.3002868
- Huang CY#, Huang YS#, Sugihara Y, Wang HY, Huang LT, Lopez-Agudelo JC, Chen YF, Lin KY, Chiang BJ,Toghani AA, Kourelis J, Wang CH, Derevnina L, Wu CH*. 2024. Subfunctionalization of NRC3 altered the genetic structure of the NicotianaNRC network. PLOS Genetics https://doi.org/10.1371/journal.pgen.1011402 (#Equal contribution, *Corresponding author)
- Chiang BJ#, Lin KY#, Chen YF, Huang CY, Goh FJ, Huang LT, Chen LH, Wu CH*. 2024. Development of a tightly regulated copper-inducible transient gene expression system in Nicotiana benthamianaincorporating suicide exon and Cre recombinase. New Phytologist https://doi.org/10.1111/nph.20021 (#Equal contribution, *Corresponding author)
- Goh FJ, Huang CY, Derevnina L, Wu CH*. 2024. NRC immune receptor networks show diversified hierarchical genetic architecture across plant lineages. The Plant Cell https://doi.org/10.1093/plcell/koae179 (*Corresponding author)
- Sakai T, Contreras MP, Martinez-Anaya C, Lüdke D, Kamoun S*, Wu CH*, Adachi H*. 2024. The NRC0 gene cluster of sensor and helper NLR immune receptors is functionally conserved across asterid plants. The Plant Cell https://doi.org/10.1093/plcell/koae154 (*Corresponding author)
- Chen JY, Sang H, Chilvers MI, Wu CH, Chang HX. 2024. Characterization of soybean chitinase genes induced by Rhizobacteria involved in the defense against Fusarium oxysporum. Plant Sci.doi: 10.3389/fpls.2024.1341181
- Wu CH and Derevnina L. 2023. The battle within: How pathogen effectors suppress NLR-mediated immunity. Current Opinion in Plant Biology org/10.1016/j.pbi.2023.102396
- Sheikh AH, Zacharia I, Pardal1 AJ, Dominguez-Ferreras A, Sueldo DJ, Kim JG, Balmuth A, Gutierrez JR, Conlan BF, Ullah N, Nippe OM, Girija AM, Wu CH, Sessa G, Jones AME, Grant MR, Gifford ML, Mudgett MB, Rathjen JP and Ntoukakis V. 2023. Dynamic changes of the Prf/Pto tomato resistance complex following effector recognition. Nature Communications 14: 2568. org/10.1038/s41467-023-38103-6
- Contreras MP, Pai Hsuan, Selvaraj M, Toghani AA, Lawson DM, Tumtas Y, Duggan C, Yuen ELH, Stevenson CEM, Harant A, Wu CH, Bozkurt TO, Kamoun S, Derevnina L. Resurrection of plant disease resistance proteins via helper NLR bioengineering. 2023. Science Advances 9, eadg3861. DOI: 10.1126/sciadv.adg3861
- Oh S, Kim S, Park HJ, Kim MS, Seo MK, Wu CH, Lee HA, Kim HS, Kamoun S, Choi D. 2023. Nucleotide-binding leucine-rich repeat network underlies nonhost resistance of pepper against the Irish potato famine pathogen Phytophthora infestans. Plant Biotechnology Journal org/10.1111/pbi.14039
- Adachi H, Sakai T, Harant A, Duggan C, Bozkurt TO, Wu CH#, Kamoun S#. 2023. An atypical NLR protein modulates the NRC immune receptor network. PLOS Genetics 19(1): e1010500. org/10.1371/journal.pgen.1010500(#Corresponding author)
- Ahn HK, Lin X, Olave-Achury AC, Derevnina L, Contreras MP, Kourelis J, Wu CH, Kamoun S, Jones JDG. 2023. Effector-dependent activation and oligomerization of plant NRC class helper NLRs by sensor NLR immune receptors Rpi-amr3 and Rpi-amr1. EMBO J e111484. org/10.15252/embj.2022111484
- Contreras MP, Pai H, Tumtas Y, Duggan C, Yuen ELH, Cruces AV, Kourelis J, Ahn HK, Lee KT, Wu CH, Bozkurt TO, Derevnina L, Kamoun S. 2023. Sensor NLR immune proteins activate oligomerization of their NRC helper. EMBO J doi.org/10.15252/embj.2022111519
- Kourelis J, Contreras MP, Harant A, Pai H, Lüdke D, Adachi H, Derevnina L, Wu CH#, Kamoun S#. 2022. The helper NLR immune protein NRC3 mediates the hypersensitive cell death caused by the cell-surface receptor Cf-4. PLOS Genetics 18(9): e1010414. org/10.1371/journal.pgen.1010414 (#Corresponding author)
- Lin X, Olave-Achury A, Heal R, Witek K, Karki HS, Song T, Wu CH, Adachi H, Kamoun S, Vleeshouwers VGAA, Jones JDG. 2022. A potato late blight resistance gene protects against multiple Phytophthora species by recognizing a broadly conserved RXLR-WY effector. Molecular Plant 15(9): 1457-1469. org/10.1016/j.molp.2022.07.012
- Derevnina L, Contreras MP, Adachi H, Upson JL, Cruces AV, Xie R, Sklenar J, Menke FLH, Mugford ST, MacLean D, Ma W, Hogenhout S, Goverse A, Maqbool A, Wu CH#, and Kamoun S#. 2021. Plant pathogens convergently evolved to counteract redundant nodes of an NLR immune receptor network. PLOS Biology 19(8): e3001136. org/10.1371/journal.pbio.3001136 (#Corresponding author)
- Duggan C, Moratto E, Savage Z, Hamilton E, Adachi H, Wu CH, Leary AY, Tumtas Y, Maqbool A, Kamoun S, Bozkurt TO. 2021. Dynamic accumulation of a helper NLR at the plant-pathogen interface underpins pathogen recognition. PNAS 118:e2104997118 org/10.1073/pnas.2104997118
- Duxbury Z, Wu CH#, Ding P#. 2021. A comparative overview of the intracellular guardians of plants and animals: NLRs in innate immunity and beyond. Rev. Plant Biol 72:155-184 doi.org/10.1146/annurev-arplant-080620-104948 (#Corresponding author)
- Witek K, Lin X, Karki HS, Jupe F, Witek AI, Steuernagel B, Stam R, van Oosterhout C, Fairhead S, Heal R, Cocker JM, Barrett W, Wu CH, Adachi H, Song T, Kamoun S, Vleeshouwers VGAA, Tomlinson L, Wulff BBH, Jones JDG. 2021. A complex resistance locus in Solanum americanum recognizes a conserved Phytophthora Nature Plants 7: 198–208. doi.org/10.1038/s41477-021-00854-9
- Wu CH and Kamoun S. 2021. Tomato Prf requires NLR helpers NRC2 and NRC3 to confer resistance against the bacterial speck pathogen Pseudomonas syringae tomato. Acta Hortic. 1316, 61-66 doi.org/10.17660/ActaHortic.2021.1316.9 bioRxiv: doi.org/10.1101/595744
- Gao C, Xu H, Huang J, Sun B, Zhang F, Savage Z, Duggan C, Yan T, Wu CH, Wang Y, Vleeshouwers VGAA, Kamoun S, Bozkurt TO, Dong S. 2020. Pathogen manipulation of chloroplast function triggers a light-dependent immune recognition. PNAS 117:9613-9620 org/10.1073/pnas.2002759117
- Wu CH*, Adachi H*, De la Concepcion JC*, Castells-Graells R, Nekrasov V, Kamoun S. 2020. NRC4 gene cluster is not essential for bacterial flagellin-triggered immunity. Plant Physiology 182: 455–459. org/10.1104/pp.19.00859 (* Equal contribution)
- Frantzeskakis L, Pietro A Di, Rep M, Schirawski J, Wu CH, and Panstruga R. 2020. Rapid evolution in plant–microbe interactions–a molecular genomics perspective. New Phytologist 225 :1134-1142. org/10.1111/nph.15966

Wen-Chi Hu (胡文綺)
Ph.D. Plant Pathology and Microbiology, National Taiwan University
wenchi6276@gate.sinica.edu.tw
+886-2-27871142 (R422)

Liang-Yu Hou (侯良諭)
Ph.D. Biology, Ludwig-Maximilians-Universität München, Germany
a090716@gate.sinica.edu.tw
+886-2-27871142 (R420)

Hung-Yu Wang (汪紘宇)
National Chung-Hsing University (TIGP-MBAS)
M.S. Bioscience and Biotechnology, National Taiwan Ocean University
hank99911@gmail.com
+886-2-27871142 (R422)

Juan Carlos Lopez (洛胡安)
National Chung-Hsing University (TIGP-MBAS)
M.S. Molecular Genetics and Biotechnology, University of Seville, Spain
as0200314@gate.sinica.edu.tw
+886-2-27871142 (R420)

Sopio Tchabashvili (莎平歐)
National Chung-Hsing University (TIGP-MBAS)
M.S. Agriculture Sciences, Georgian Technical University, Georgia
tchabashvil0001@gate.sinica.edu.tw
+886-2-27871142 (R420)

Kuan-Yu Lin (林冠妤)
National Taiwan University
b08608046@ntu.edu.tw
+886-2-27871142 (R422)

Chin-Wen Chang (張槿玟)
M.S. Plant Biology, National Taiwan University
jwchang@gate.sinica.edu.tw
+886-2-27871142 (R420)

Bing-Jen Chiang (江秉真)
M.S. Plant Pathology and Microbiology, National Taiwan University
as0191504@gate.sinica.edu.tw
+886-2-27871142 (R420)

Yi-Feng Chen (陳宜豐)
M.S. Plant Pathology and Microbiology, National Taiwan University
yfchen00@gate.sinica.edu.tw
+886-2-27871142 (R420)

Wen-Rong Hsiao (蕭文榮)
M.S. Plant Pathology, National Chung-Hsing University
wrhsiao@gate.sinica.edu.tw
+886-2-27871142 (R420)
國際
- 2024 歐洲分子生物學組織(EMBO)全球研究學者 - 歐洲分子生物學組織 (EMBO)
國內
- 2025 前瞻計畫 - 中央研究院
- 2025 年輕學者創新獎 - 財團法人傑出人才發展基金會
- 2024 楊祥發院士傑出農業科學年輕學者獎 - 財團法人楊祥發紀念教育基金會
- 2024 新秀獎 - 臺灣植物學會
- 2022 2030跨世代年輕學者–新秀學者研究計畫 - 科技部