[Pao-Yang Chen] Estimating genome-wide DNA methylation heterogeneity with methylation patterns
POST: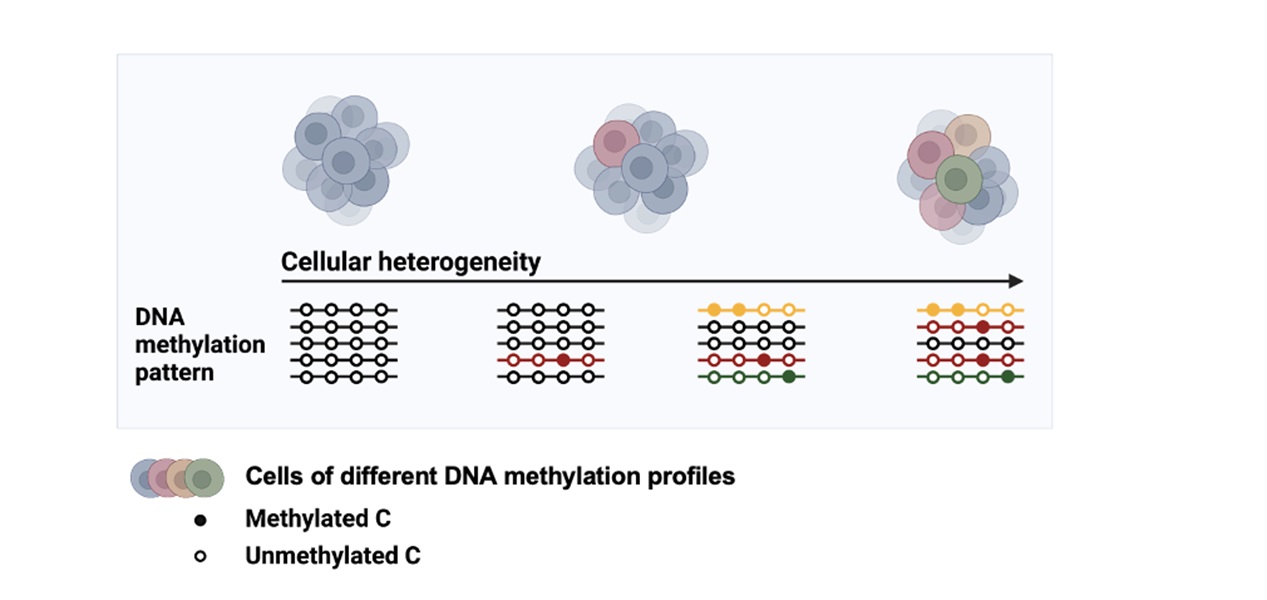
Figure. Illustrations of the DNA methylation patterns as a proxy for monitoring cellular development.
In a heterogeneous population of cells, individual cells can behave differently and respond variably to the environment. This cellular diversity can be assessed by measuring DNA methylation patterns. The loci with variable methylation patterns are informative of cellular heterogeneity and may serve as biomarkers of diseases and developmental progression. Cell-to-cell methylation heterogeneity can be evaluated through single-cell methylomes or computational techniques for pooled cells. However, the feasibility and performance of these approaches to precisely estimate methylation heterogeneity require further assessment. Here, we proposed model-based methods adopted from a mathematical framework originally from biodiversity, to estimate genome-wide DNA methylation heterogeneity. We evaluated the performance of our models and the existing methods with feature comparison, and tested on both synthetic datasets and real data. Overall, our methods have demonstrated advantages over others because of their better correlation with the actual heterogeneity. We also demonstrated that methylation heterogeneity offers an additional layer of biological information distinct from the conventional methylation level. In the case studies, we showed that distinct profiles of methylation heterogeneity in CG and non-CG methylation can predict the regulatory roles between genomic elements in Arabidopsis. This opens up a new direction for plant epigenomics. Finally, we demonstrated that our score might be able to identify loci in human cancer samples as putative biomarkers for early cancer detection. We adopted the mathematical framework from biodiversity into three model-based methods for analysing genome-wide DNA methylation heterogeneity in order to monitor cellular heterogeneity. Our methods, namely MeH (https://github.com/PaoyangLab/MeH), have been implemented, evaluated with existing methods, and are open to the research community.