Dynamic Localization of Spo11-1 and Conformational Changes of Meiotic Axial Elements During Recombination Initiation of Maize Meiosis
Jia-Chi Ku, Arnaud Ronceret, Inna Golubovskaya, Ding Hua Lee, Chiting Wang, Ljudmilla Timofejeva, Yu-Hsin Kao, Ana Angoa, Karl Kremling, Rosalind Williams-Carrier, Robert Meeley, Alice Barkan, W. Zacheus Cande and Chung-Ju Rachel Wang
Meiotic recombination is essential for genetic diversity, which is initiated with DNA double-strand breaks (DSBs) catalyzed by the topoisomerase-related enzyme, SPO11. The activity, timing and location of this DSB machinery must be controlled precisely, but how this is achieved remains obscure. The research team led by Dr. Chung-Ju Wang showed dynamic localization of thousands of SPO11-1 on chromatin during meiotic initiation in maize, yet a similar number of SPO11-1 corresponding to the number of DSBs is able to load onto axial elements (AEs), which accompanies a structural change of the AEs of wild-type meiotic chromosomes. Interestingly, loss of SPO11-1 not only affects DSB formation but also impairs structural alterations of AEs, resulting in abnormally long and curly AEs during early meiosis. This study provides the first cytological observation in all organisms for dynamic distribution of SPO11-1 during recombination initiation and suggests an intimate relationship between DSB formation and AE structural changes. This work was selected as the PLOS Genetics journal cover.
PLOS Genetics 16(4), e1007881 (2020).
DOI: 10.1371/journal.pgen.1007881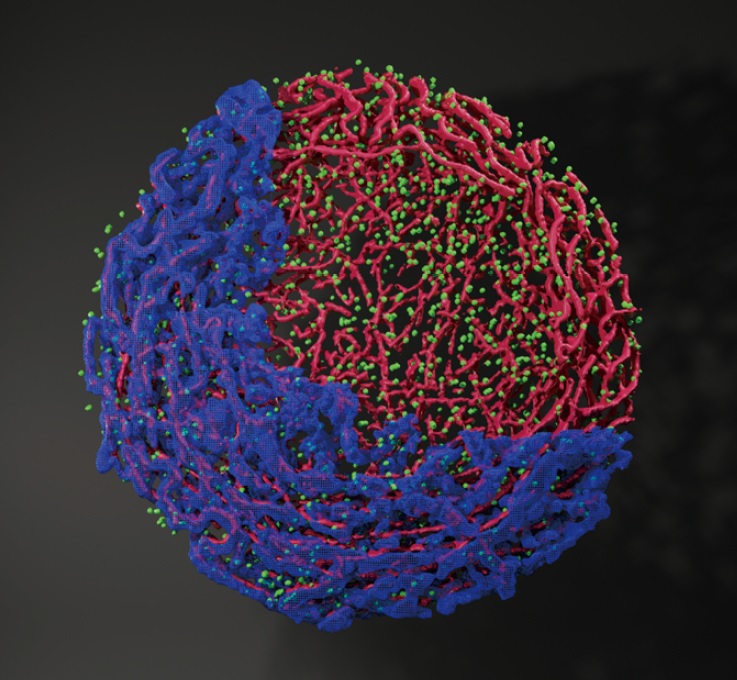
A 3D surface rendering image of a maize meiotic nucleus via super-resolution microscopy showed the first cytological observation in all organisms for the distribution relationship among SPO11 (green), chromosome axes (red) and chromatin (blue). To manifest their relative positions, chromatin is partially displayed.