Jauh, Guang-Yuh (趙光裕)
Deputy Director, Administrative Affairs
Research Fellow
- Ph.D., Botany and Plant Sciences, University of California, Riverside, USA
- The molecular and cellular mechanisms of genes involved in pollen tube elongation, embryogenesis, and seed development.
- jauh@gate.sinica.edu.tw
- jauh@as.edu.tw
- +886-2-2787-1024 (Lab: R126)
- +886-2-2787-1156 (Office: R124)
- Academia Sinica Archive
- ORCID
- Web of Science (WOS)
- Google Scholar
Seeds and grains constitute the major source of food supply around the world, and their production depends on successful double fertilization and subsequent embryogenesis. To meet the needs of the fast-growing population and dramatic global weather change, the yield potential of crops must be increased. With my prime expertise in plant cell biology, my long-term research interests focus on exploring the biological functions of genes/proteins in plant sexual reproduction, including microspore development, pollen tube elongation, pollination, and embryogenesis. Few years ago, we initiated a bioinformatic approach to search the promising T-DNA insertional mutants with defective gametophytic and/or embryogenetic development in all Arabidopsis genes. After careful phenotyping, genotyping and complementation with their genomic sequences, more than one dozen of uncharacterized genes were obtained and some of them were chosen for functional study. These T-DNA-insertional mutants showing severe defects during plant reproduction and lacking homozygote progenies were called no homozygote progeny mutants, nhps.
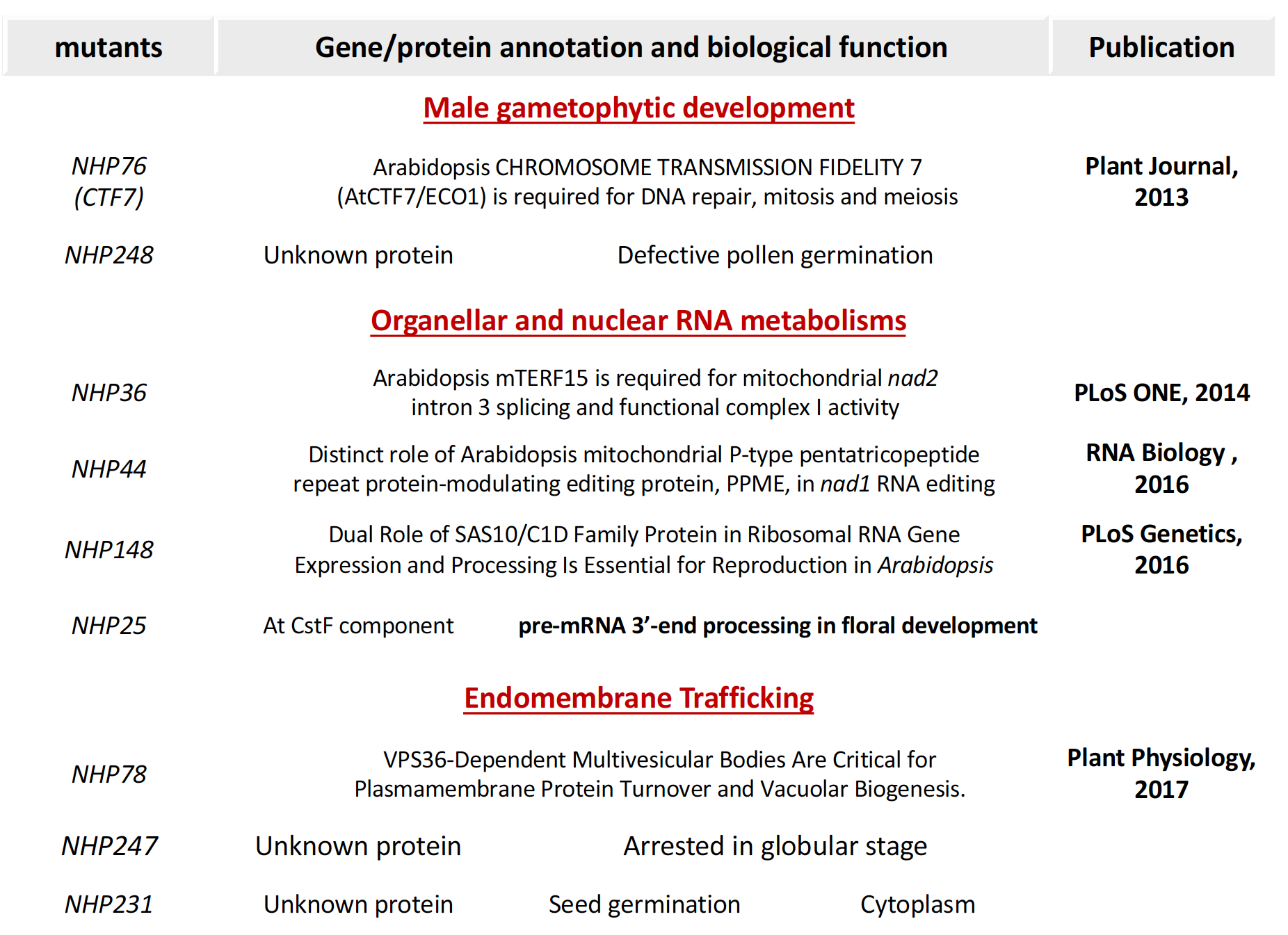

Based on the cellular functions of encoding proteins, they are classified into two categories: RNA metabolisms and unique cellular processes. For the RNA metabolisms, NHP36 and NHP44 encode an Arabidopsis mTERF15 (Hsu et al., 2014) and a P type PPR proteins (Leu et al., 2016) required for mitochondrial transcripts splicing and editing, respectively. NHP148 encodes an Arabidopsis nucleolus-localized SAS10/C1D Family Protein required for rRNA post-transcriptional processes (Chen et al., 2016). For the unique cellular processes, NHP78 is a VPS36 required for late endosome formation and targeting ubiquitylated plasma membrane protein to vacuole/lysosome degradation (Wang et al., 2017). Currently, the molecular and cellular mechanisms of several NHP genes are under investigated. Our preliminary results show their unique roles in floral development (NHP25), cytokinesis during embryogenesis (NHP248), and GA mediated seed germination (NHP231).
In flowering plants, pollen, the male gametophyte, deposited on the stigma of a flower germinates by producing a pollen tube that grows through the style and into the micropyle of an ovule to release its two sperm cells for double fertilization with the egg and central cell. Understanding the pollen tube gene functions involved in gametophytic development and pollination will provide new insights into the complicated regulatory machinery involved in these crucial processes. Nevertheless, it has become increasingly clear that signals from the maternal tissue are vital for in vivo processes. A number of pollen-specific genes have been described closely with respect to the timing of transcription and translation during pollen maturation, germination and tube cell growth. Nevertheless, most of these studies were conducted under in vitro conditions due to the difficulty in collecting enough in vivo pollen tubes within the solid maternal gynoecium. By using a pollen-specific promoter (ProLAT52) to generate an epitope-tagged polysomal-RNA complex, we obtained mRNAs undergoing translation (the translatome) of in vivo-grown pollen tubes from self-pollinated Arabidopsis gynoecia. Translatomes of pollen grains as well as in vivo and in vitro cultured pollen tubes were assayed by microarray analyses, which revealed more than 500 transcripts specifically enriched in in vivo elongating pollen tubes. Functional analyses of several in vivo mutants of these pollination-enhanced transcripts revealed partial pollination, fertilization and seed formation defects in siliques (Lin et al., 2014). Cytology observation confirmed the involvement of these genes in specialized processes including micropylar guidance, pollen tube burst and repulsion of multiple pollen tubes in embryo sacs. For example, two Arabidopsis Small Auxin Up RNA genes, SAUR62 and SAUR75, show expression upregulated by pollination. Both SAUR62 and SAUR75 mainly localized in pollen tube nuclei, and siliques of homozygous saur62 (saur62/-), saur75 (saur75/-), and the SAUR62/75 RNAi knockdown line had many aborted seeds. Pollen viability of these mutants and RNAi lines was normal, but in vitro and in vivo pollen tube growth was defective, with branching
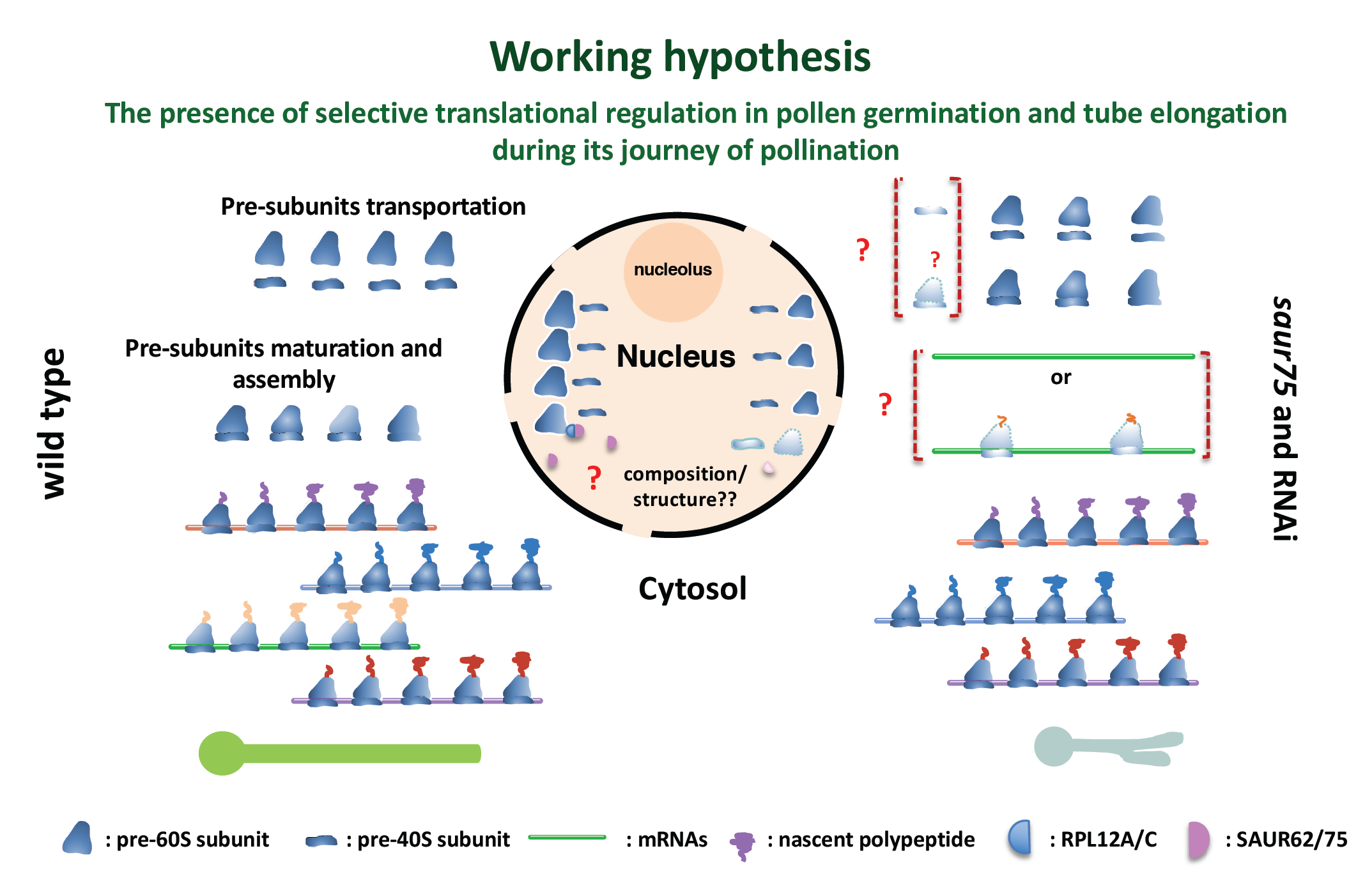
phenotypes. Immunoprecipitation with transgenic SAUR62/75-GFP flowers revealed ribosomal protein RPL12 family members as their partners. Polysome profiling showed reduced 80S ribosome abundance in homozygous saur62, saur75, rpl12c and SAUR62/75 RNAi flowers, which suggests roles for SAUR62/75 in ribosome assembly. To clarify the roles in translation, total proteins of wild-type (WT) and RNAi flowers were analyzed by iTRAQ, which revealed significantly reduced expression of factors participating in pollen tube wall biogenesis and F-actin dynamics, critical for the elastic properties of tube elongation. Indeed, RNAi pollen tubes showed dislocalization of de-esterified and esterified pectins and F-actin organization. Here, we demonstrate the biological roles of SAUR62/75 and their partners, RPL12 family members, as critical in ribosome assembly for efficient pollen tube elongation and the following fertilization (He et al., 2018).
- Hsu YW, Juan CT, Guo CL, Wang HJ, Jauh GY*. (2025), HYCCIN2 is essential for recruiting the SH3P2-DRP1A complex to the cell plate in Arabidopsis embryos. New Phytol, 247: 2781. https://doi.org/10.1111/nph.70367
- Yen C-C, Hsu Y-W, Leu K-C, Chen S-S, Chen T-Y, Juan C-T, Wang C-, Kuan C, Shaw J-F, Ho C-M K*, Jauh G-Y*. Guard-cell phytosterol homeostasis is critical for proper stomatal patterning. bioRxiv, 2024. 08.30. doi: https://doi.org/10.1101/2024.08.29.610202.
- Yen CC, Hsu CM, Jiang PL, Jauh GY*. (2024). Dynamic organelle changes and autophagic processes in lily pollen germination. Bot Stud. 65: 5.
- Radjacommare R, Lin SY, Usharani R, Lin WD, Jauh GY, Schmidt W, Fu H. * (2023). The Arabidopsis deubiquitylase OTU5 suppresses flowering by histone modification-mediated activation of the major flowering repressors FLC, MAF4, and MAF5. Int J Mol Sci. 24:6176.
- Hsu PJ, Tan MC, Shen HL, Chen YH, Wang YY, Hwang SG, Chiang MH, Le QV, Kuo WS, Chou YC, Lin SY, Jauh GY, Cheng WH*. (2021) The nucleolar protein SAHY1 is involved in pre-rRNA processing and normal plant growth. Plant Physiology 185:1039-1058.
- Hsu YW, Juan CT, Jauh GY* (2019) Mitochondrial HSP60s interact with WTF9 to regulate RNA splicing of ccmFC and rpl2. Plant Cell Physiol. 60(1):116-125.
- He SL, Hsieh HL, Jauh GY* (2018) Mutation of Arabidopsis SAURsimpairs the efficient translation of transcripts essential for pollen tube growth. Plant Physiology (in press)
- Hsu Y-W, Jauh GY* (2017). VPS36-Mediated Plasma Membrane Protein Turnover is Critical for Arabidopsis Root Gravitropism. Plant Signal Behav. doi10.1080/15592324. 2017.1307495
- Huei-Jing Wang, Ya-Wen Hsu, Cian-Ling Guo, Wann-Neng Jane, Hao Wang, Liwen Jiang and Jauh GY* (2017). VPS36-dependent Multivesicular Bodies are Critical for Plasmamembrane Protein Turnover and Vacuolar Biogenesis. Plant Physiology 173: 566-581.
- Chen YJC, Wang HJ, Jauh GY* (2016). Dual role of a SAS10/C1D family Protein in ribosomal RNA gene expression and processing is essential for reproduction in Arabidopsis thaliana. PLoS Genet 12(10): e1006408.
- Leu KC, Hsieh MH, Wang HJ, Hsieh HL, Jauh GY* (2016). Distinct role for the Arabidopsis mitochondrial P-type pentatricopeptide repeat protein modulating editing protein, PPME, in nad1 RNA editing. RNA Biology 3:593-604
- Bolaños-Villegas P*, Jauh GY (2015) Reduced activity of Arabidopsis chromosome- cohesion regulator gene CTF7/ECO1 alters cytosine methylation status and retrotransposon expression. Plant Signaling & Behavior 10(5): e1013794.
- Hsu YW, Wang HJ, Hsieh MH, Hsieh HL, Jauh GY* (2014) Arabidopsis mTERF15 is required for mitochondrial nad2 Intron 3 splicing and functional complex I activity. PLoS ONE. 9(11): e112360.
- Lin SY, Chen PW, Chuang MH, Juntawong P, Bailey-Serres J, Jauh GY* (2014) Profiling of Translatomes of in Vivo-Grown Pollen Tubes Reveals Genes with Roles in Micropylar Guidance during Pollination in Arabidopsis. The Plant Cell. 26: 602-618 (highlighted in the “IN BRIEF” of the same issue).
- Bolaños-Villegas P, Yang X, Wang HJ, Juan CT, Chuang MH, Makaroff CA, Jauh GY* (2013) Arabidopsis CHROMOSOME TRANSMISSION FIDELITY 7 (AtCTF7/ECO1) is required for DNA repair, mitosis and meiosis. Plant J. 75: 927-940.
Domestic
- 2000: Li Foundation prize, Li Foundation
This PI participates in the following graduate programs.
