[Chen, Pao-Yang] Computational Methods for Assessing Chromatin Hierarchy
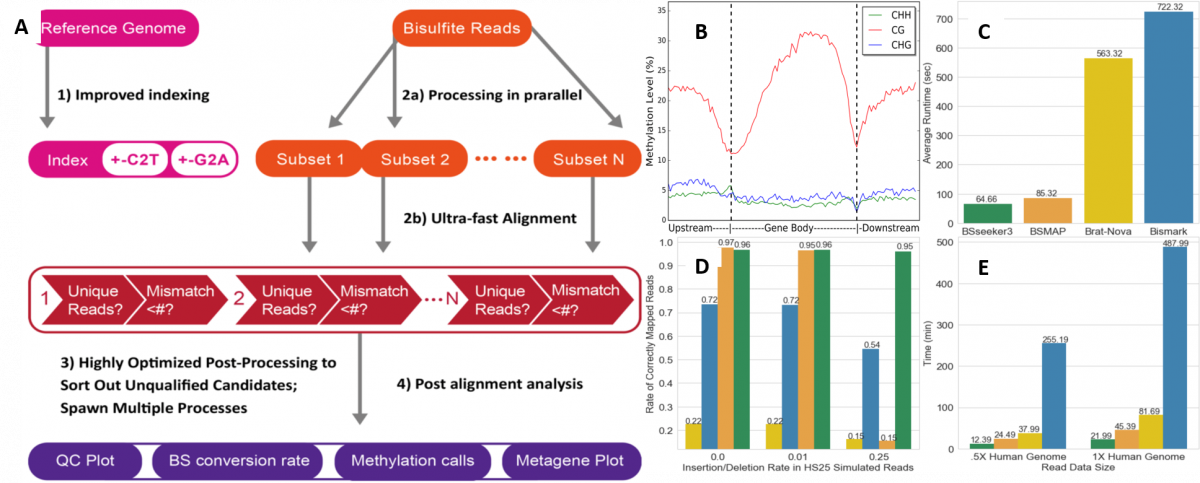
POST:Summary of BS-Seeker3 pipeline and performance. (A) Schematic flow chart of BS Seeker 3 with improved indexing, data processing, fast alignment and post-alignment analyses (B) Metaplot of Methylation level: This metaplot presents the average methylation level distribution within a user-specified genomic structure (e.g., coding genes) in Arabiodopsis thalania. CG denotes a CpG dinucleotide, CHG denotes a cytosine next to a H where H stands for A, C, or T.and then a guanine, CHH denotes a cytosine next to two H bases (C) Average user runtime of the four aligners on 10M simulated HiSeq 2500 Arabidopsis reads. (D) Percentage of the 10M simulated HiSeq2500 reads that were mapped correctly across various reads complexity level. (E) Average runtime of four aligners on directional BS-seq reads from real human data.
Background: DNA methylation is an important epigenetic modification critical in regulation and transgenerational inheritance. The methylation level can be estimated at single-nucleotide resolution by whole-genome bisulfite sequencing (BS-seq; WGBS). Current bisulfite aligners provide pipelines for processing the reads by WGBS; however, few are able to analyze the BS-seqs in a reasonable timeframe that meets the needs of the rapid expansion of epigenome sequencing in biomedical research.
Results: We introduce BS-Seeker3, an extensively improved and optimized implementation of BS-Seeker2 that leverages the available computational power of a standard bioinformatics lab.
BS-Seeker3 adopts all alignment features of BS-Seeker2. It performs ultrafast alignments and achieves both high accuracy and high mappability, more than twice that of the other aligners that we evaluated. Moreover, BS Seeker 3 is well linked with downstream analyzer MethGo for up to 9 types of genomic and epigenomic analyses. BS-Seeker3 is an accurate, versatile, ultra-fast pipeline for processing bisulfite-converted reads. It also helps the user better visualize the methylation data.
Availability: BS Seeker3 and a tutorial are freely available at https://github.com/khuang28jhu/bs3/