[Pao-Yang Chen] Diversity of fungal DNA methyltransferases and their association with DNA methylation patterns
POST: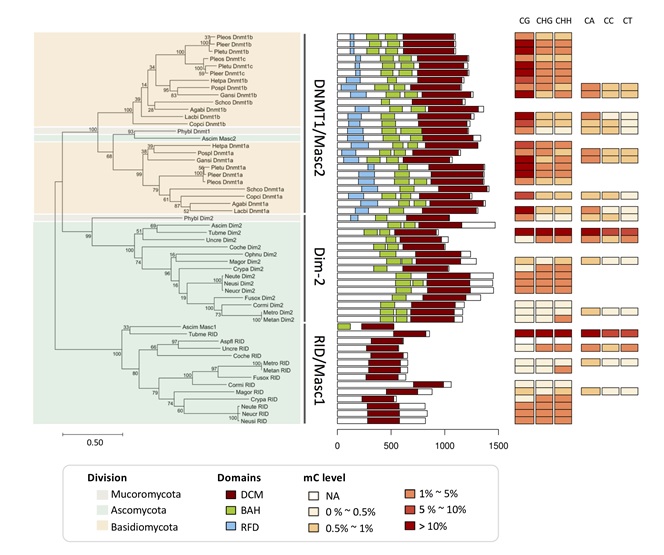
Figure: Fungal phylogenetic and domain architecture analyses of the DNMT1 family and RID/Masc1 protein together with the fungal DNA methylation level.
DNA methyltransferases (DNMTs) are a group of proteins that catalyze DNA methylation by transferring a methyl group to DNA. The genetic variation in DNMTs results in differential DNA methylation patterns associated with various biological processes. In fungal species, DNMTs and their DNA methylation profiles were found to be very diverse and have gained many research interests. We reviewed fungal DNMTs in terms of their biological functions, protein domain structures, and their associated epigenetic regulations compared to those known in plant and animal systems. In addition, we summarized recent reports on potential RNA-directed DNA methylation (RdDM) related to DNMT5 in fungi. We surveyed up to 40 fungal species with published genome-wide DNA methylation profiles (methylomes) and presented the associations between the specific patterns of fungal DNA methylation and their DNMTs based on a phylogenetic tree of protein domain structures. For example, the main DNMTs in Basidiomycota, DNMT1 with RFD domain +DNMT5, contributing to CG methylation preference, were distinct from RID+Dim-2 in Ascomycota, resulting in a non-CG methylation preference. Lastly, we revealed that the dynamic methylation involved in fungal life stage changes was particularly low in mycelium and DNA methylation was preferentially located in transposable elements (TEs). This review comprehensively discussed fungal DNMTs and methylomes and their connection with fungal development and taxonomy to present the diverse usages of DNA methylation in fungal genomes.
The first authors of this study are Dr. Yu-Shin Nai of Department of Entomology, National Chung Hsing University, and the PhD student Yu-Chun Huang from TIGP-Bioinformatics Program.