[Chuan Ku] Host range and coding potential of eukaryotic giant viruses
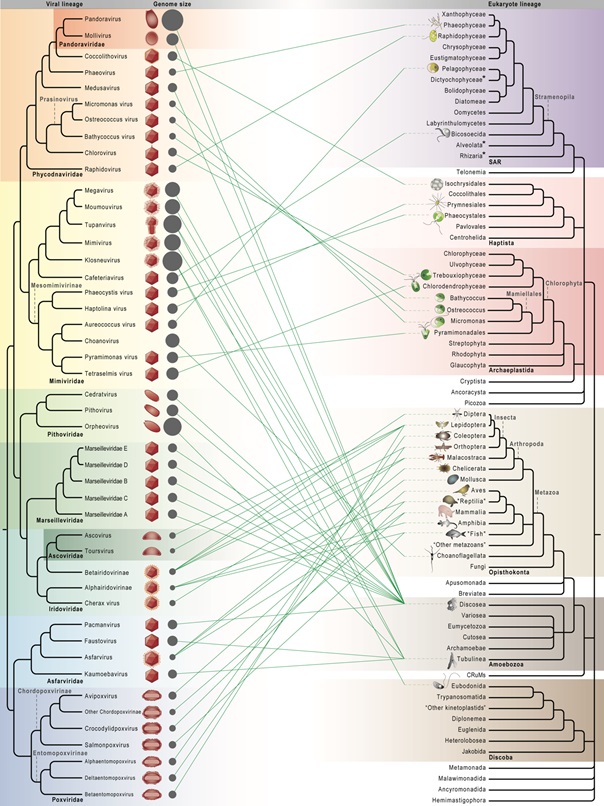
POST:Giant viruses are a group of eukaryotic double-stranded DNA viruses with large virion and genome size that challenged the traditional view of virus. Since 2001, when four diverse viral families were suggested to form a monophyletic group called Nucleo-Cytoplasmic Large DNA Viruses (now phylum Nucleocytoviricota), newly isolated strains and sequenced genomes have substantially advanced our knowledge of their host diversity, gene functions, and evolutionary history. Giant viruses are now known to infect hosts from all major supergroups in the eukaryotic tree of life, which predominantly comprises microbial organisms. The seven well-recognized viral clades (taxonomic families) have drastically different host range. Mimiviridae and Phycodnaviridae, both with notable intrafamilial genome variation and high abundance in environmental samples, are no longer limited to amoebal and algal hosts, from which they were originally identified, and are now found to infect multiple eukaryotic supergroups. Laboratory experiments and comparative genomics have shed light on the unprecedented functional potential of giant viruses, encoding proteins for genetic information flow, energy metabolism, synthesis of biomolecules, membrane transport, and sensing that allow for sophisticated control of intracellular conditions and cell-environment interactions. Evolutionary genomics can illuminate how current and past hosts shape viral gene repertoires, although it becomes more obscure with divergent sequences and deep phylogenies. Continued works to characterize giant viruses from marine and other environments will further contribute to our understanding of their host range, coding potential, and virus-host coevolution.