[Yue-Ie Hsing] Genomes of 13 domesticated and wild rice relatives highlight genetic conservation, turnover and innovation across the genus Oryza (cover story of Nature Genetics at Feb 2018)
POST:
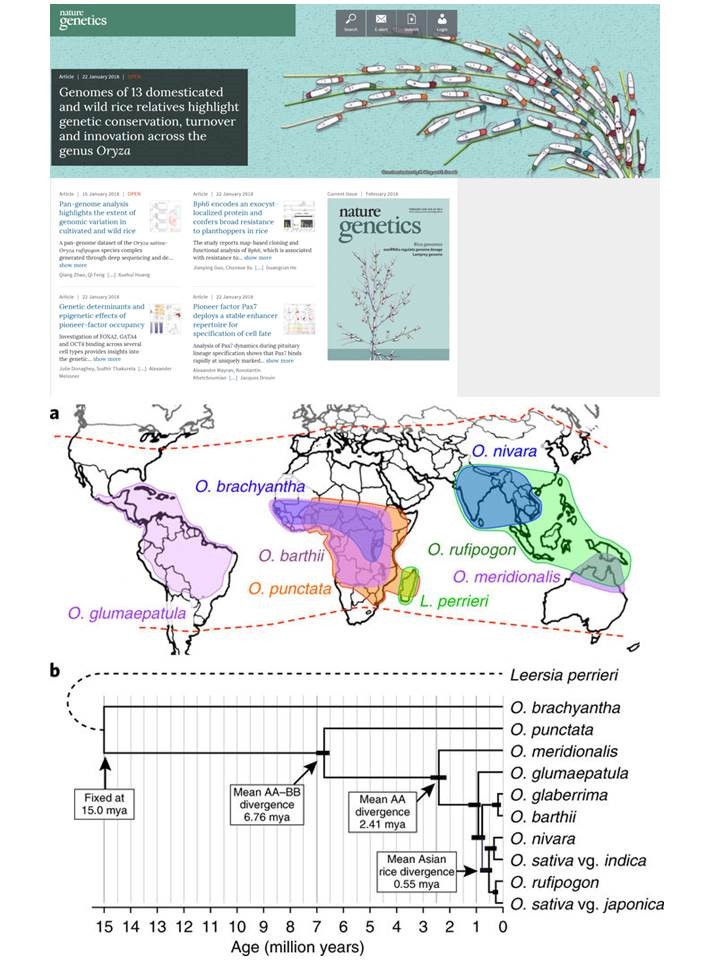
The genus Oryza is a model system for the study of molecular evolution over time scales ranging from a few thousand to 15 million years. Using 13 reference genomes spanning the Oryza species tree, we show that despite few large-scale chromosomal rearrangements rapid species diversification is mirrored by lineage-specific emergence and turnover of many novel elements, including transposons, and potential new coding and noncoding genes. Our study resolves controversial areas of the Oryza phylogeny, showing a complex history of introgression among different chromosomes in the young ‘AA’ subclade containing the two domesticated species. This study highlights the prevalence of functionally coupled disease resistance genes and identifies many new haplotypes of potential use for future crop protection. Finally, this study marks a milestone in modern rice research with the release of a complete long-read assembly of IR 8 ‘Miracle Rice’, which relieved famine and drove the Green Revolution in Asia 50 years ago. This is one of the main papers published from the International Oryza map alignment project initiated at 2007. More than 50 scientists joined this work and 5 of them are from our Institute. This work was selected as cover story of February issue of Nature Genetics.This work was the cover story of the February 2018 issue of Nature Genetics. The figures illustrate the distribution of the Oryza species we sequenced, and their phylogenic relationship.