Modular evolution of secretion systems and virulence plasmids in a bacterial species complex
Lin Chou, Yu-Chen Lin, Mindia Haryono, Mary Nia M. Santos, Shu-Ting Cho, Alexandra J. Weisberg, Chih-Feng Wu, Jeff H. Chang, Erh-Min Lai and Chih-Horng Kuo
Many named species in bacterial taxonomy correspond to species complexes, in which the species boundaries and inter-species relationships are uncertain. Such uncertainties hamper research, as well as impact pathogen identification and policy-making for quarantine regulations. To solve this challenge, a team led by Dr. Chih-Horng Kuo at Institute of Plant and Microbial Biology, Academia Sinica, utilized the Agrobacterium tumefaciens species complex as the study system, devised strategies for genomic analysis, and investigated the evolution of secretion systems and virulence plasmids. The findings reveal a complex history of evolution processes shaping the diversity of effectors and provide explanation on the differences in virulence against various plant hosts among these strains. The result was published in BMC Biology (January, 2022).
BMC Biology 2022, 20:16.
DOI: 10.1186/s12915-021-01221-y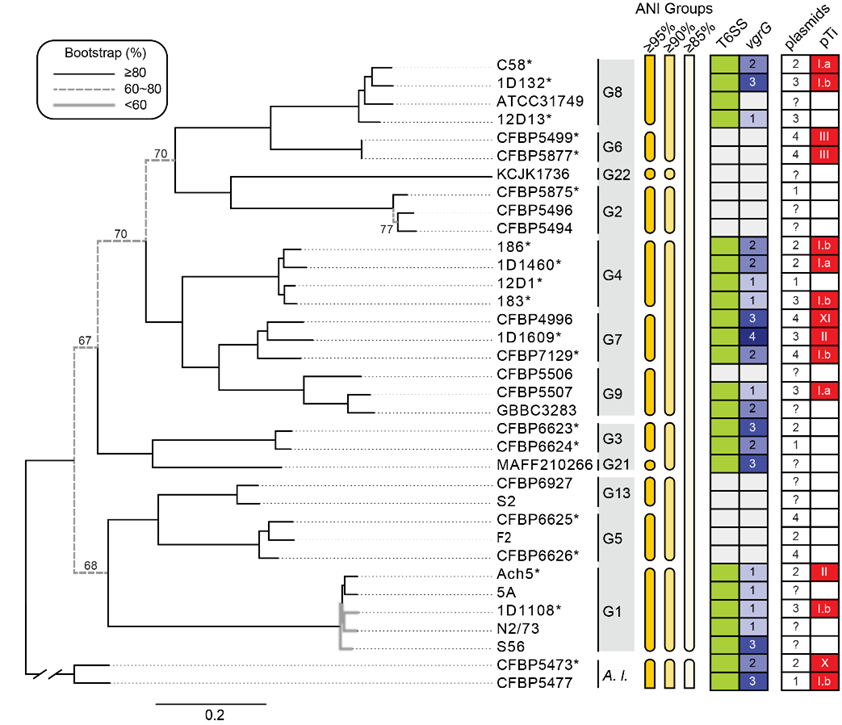
Relationships among representatives of the Agrobacterium tumefaciens species complex. The genomospecies assignments (i.e., G1-G9, G13, and G21-G22), grouping based on genome-wide average nucleotide identity (ANI), presence/absence of type VI secretion system (T6SS)-encoding gene cluster (green: present; white: absent), copy number of vgrG (white background: absent), number of plasmids, and the tumor-inducing plasmid (pTi) type (white background: absent) are labeled.