About Us
- We are a research group at the Institute of Plant and Microbial Biology, Academia Sinica, Taiwan
- Research areas: Evolution, Genomics, Transcriptomics, Bioinformatics, Taxonomy, Microbiology
- Principal investigator: Chih-Horng Kuo
研究室簡介
- 本研究室隸屬於 中央研究院 植物暨微生物學研究所
- 研究領域:演化生物學、基因體學、轉錄體學、生物資訊學、分類學、微生物學
- 研究室主持人:郭志鴻
Research: Evolutionary and Functional Genomics of Host-Associated Bacteria
Our research focuses on the genomic basis of bacterial diversity and microbial interactions. We integrate evolutionary genomics, comparative transcriptomics, phenotypic characterization, and molecular genetics to study bacteria with diverse ecological niches and pathogenic potential. Building on these strengths, we have established extensive collaborations to connect genotypic variations with phenotypic outcomes and dissect the molecular mechanisms of microbe-host interactions.
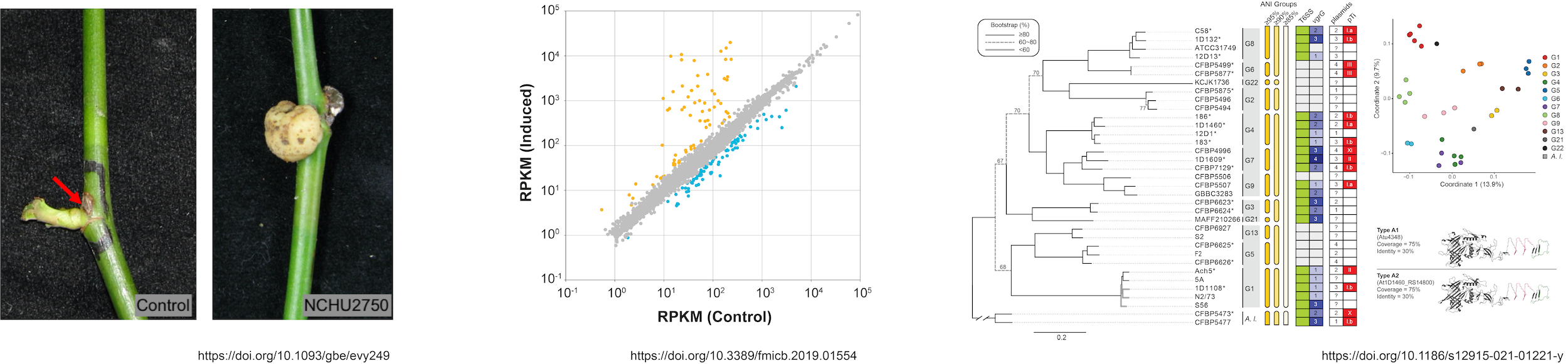
(1) Agrobacteria and Rhizobia
Agrobacteria and rhizobia are plant-associated bacteria encompassing multiple genera and spanning a wide range of ecological roles, from pathogenicity to nitrogen-fixing symbiosis. We investigate the genomic diversity and evolutionary history of this group, with a focus on the genetic basis of interactions with plants and other microbes, including the type IV secretion system (T4SS), which mediates DNA transfer into host cells, the type VI secretion system (T6SS), which contributes to interbacterial competition, and the global regulation of gene expression. These studies advance our understanding of microbe-plant and microbe-microbe interactions, and improve Agrobacterium-mediated transformation (AMT) technologies.
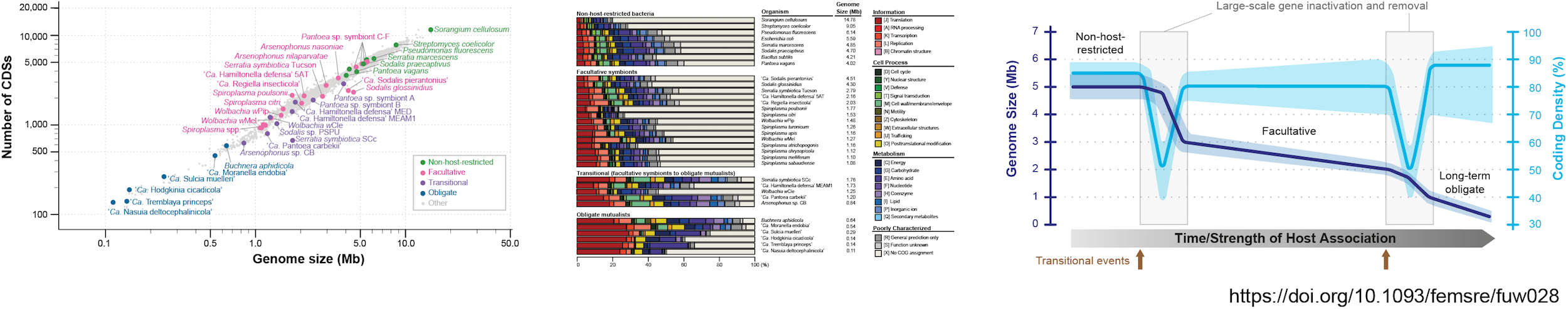
(2) Mollicutes
The class Mollicutes comprises diverse groups of host-associated bacteria, ranging from pathogens of animals and plants to protective symbionts of insects, with notable genera such as Mycoplasma, Spiroplasma, and 'Candidatus Phytoplasma'. Our studies encompass the entire class, with phytoplasmas as our primary study system. We investigate genome organization and dynamics, molecular evolution that shapes genetic divergence across lineages, roles of mobile genetic elements in horizontal gene transfer, and effectors that manipulate host development. These studies deepen our knowledge of how bacteria interact with and reprogram the development of their hosts. Our work on genome-based taxonomy of these bacteria further facilitates accurate classification and molecular diagnostics.
(3) Microbial Diversity and Comparative Genomics
Beyond our primary research systems, our expertise in comparative genomics and molecular evolution supports a broad range of collaborative studies across diverse organisms and ecological contexts. These projects extend from genomic analyses into microbiome, symbiosis, taxonomy, and applied microbiology.
研究主題:宿主相關細菌之演化與功能性基因體學
本研究室主要以基因體學探討細菌多樣性及微生物交互作用。我們整合演化基因體學、比較轉錄體學、表現型鑑定及分子遺傳學,研究具有不同生態棲位與致病潛力的細菌。奠基於這些研究方法,我們建立了廣泛的合作網絡,藉以更有效連結基因型與表型變異,並探究微生物與宿主互動的分子機制。
(1) 農桿菌與根瘤菌
農桿菌與根瘤菌為一群與植物相關的細菌,涵蓋多個屬,在生態角色上包含植物病原菌及固氮共生菌。本研究室探討這群細菌的基因體多樣性與演化歷程,著重於其與植物及其他微生物交互作用的遺傳基礎,包括介導DNA轉移至宿主細胞的第四型分泌系統、促進細菌間競爭的第六型分泌系統,以及基因表現的整體性調控。這些研究深化了我們對微生物與植物及其他微生物之間交互作用的理解,並改良農桿菌介導之基因轉殖技術。
(2) 柔膜菌綱
柔膜菌綱為一群宿主依存性細菌,生態上包含動植物病原及昆蟲之保護性共生菌等,具代表性的屬包括黴漿菌、螺旋菌質體及植物菌質體等。本團隊研究範疇涵蓋整個柔膜菌綱,尤以植物菌質體為主要研究系統。我們探討基因體組成與動態、影響不同支系遺傳分化的分子演化、移動式遺傳元件在水平基因轉移中的角色,以及操控宿主發育的效應蛋白等。這些研究加深了我們對細菌如何與宿主互動並改變宿主發育進程的知識。此外,針對基因體分類學的研究,則可改善分類準確性及分子診斷的應用。
(3) 微生物多樣性與比較基因體學
除上述主要研究系統外,本研究室基於比較基因體學與分子演化的專長,發展了多項涵蓋不同物種及生態環境的合作計畫。這些計畫奠基於基因體學分析,並延伸至微生物相、共生、分類學及應用微生物學等領域。